The clustered regularly interspaced short palindromic repeat (CRISPR) technology holds the promise to revolutionize gene editing technologies, which is transformative to the way we understand and treat diseases. This technique is based in a natural mechanism found in bacteria that allows a protein coupled to a single guide RNA (gRNA) strand to locate and make cuts in specific sites in the targeted genome. Being able to computationally predict the efficiency and specificity of gRNA is central to the success of gene editing.
Transcribed from DNA sequences, RNA is an important type of biological sequence of ribonucleotides (A, U, G, C), which folds into 3D structure. Benefiting from recent advance in large language models (LLMs), a variety of computational biology tasks can be solved by fine-tuning biological LLMs pre-trained on billions of known biological sequences. The downstream tasks on RNAs are relatively understudied.
In this post, we adopt a pre-trained genomic LLMs for gRNA efficiency prediction. The idea is to treat a computer designed gRNA as a sentence, and fine-tune the LLM to perform sentence-level regression tasks analogous to sentiment analysis. We used Parameter-Efficient Fine-Tuning methods to reduce the number of parameters and GPU usage for this task.
Solution overview
Large language models (LLMs) have gained a lot of interest for their ability to encode syntax and semantics of natural languages. The neural architecture behind LLMs are transformers, which are comprised of attention-based encoder-decoder blocks that generate an internal representation of the data they are trained from (encoder) and are able to generate sequences in the same latent space that resemble the original data (decoder). Due to their success in natural language, recent works have explored the use of LLMs for molecular biology information, which is sequential in nature.
DNABERT is a pre-trained transformer model with non-overlapping human DNA sequence data. The backbone is a BERT architecture made up of 12 encoding layers. The authors of this model report that DNABERT is able to capture a good feature representation of the human genome that enables state-of-the-art performance on downstream tasks like promoter prediction and splice/binding site identification. We decided to use this model as the foundation for our experiments.
Despite the success and popular adoption of LLMs, fine-tuning these models can be difficult because of the number of parameters and computation necessary for it. For this reason, Parameter-Efficient Fine-Tuning (PEFT) methods have been developed. In this post, we use one of these methods, called LoRA (Low-Rank Adaptation). We introduce the method in the following sections.
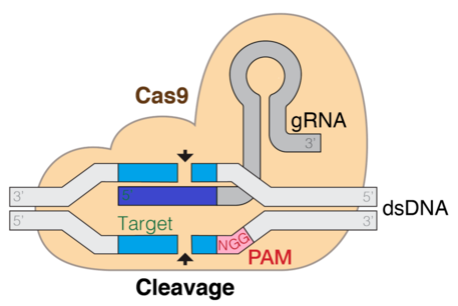
The following diagram is a representation of the Cas9 DNA target mechanism. The gRNA is the component that helps target the cleavage site.
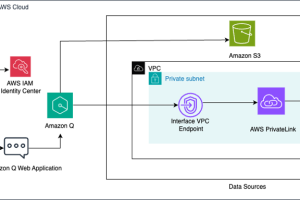
The goal of this solution is to fine-tune a base DNABERT model to predict activity efficiency from different gRNA candidates. As such, our solution first takes gRNA data and processes it, as described later in this post. Then we use an Amazon SageMaker notebook and the Hugging Face PEFT library to fine-tune the DNABERT model with the processed RNA data. The label we want to predict is the efficiency score as it was calculated in experimental conditions testing with the actual RNA sequences in cell cultures. Those scores describe a balance between being able to edit the genome and not damage DNA that wasn’t targeted.
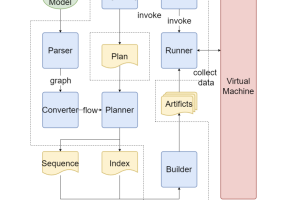
The following diagram illustrates the workflow of the proposed solution.

Prerequisites
For this solution, you need access to the following:
- A SageMaker notebook instance (we trained the model on an ml.g4dn.8xlarge instance with a single NVIDIA T4 GPU)
- transformers-4.34.1
- peft-0.5.0
- DNABERT 6
Dataset
For this post, we use the gRNA data released by researchers in a paper about gRNA prediction using deep learning. This dataset contains efficiency scores calculated for different gRNAs. In this section, we describe the process we followed to create the training and evaluation datasets for this task.
To train the model, you need a 30-mer gRNA sequence and efficiency score. A k-mer is a contiguous sequence of k nucleotide bases extracted from a longer DNA or RNA sequence. For example, if you have the DNA sequence “ATCGATCG” and you choose k = 3, then the k-mers within this sequence would be “ATC,” “TCG,” “CGA,” “GAT,” and “ATC.”
Efficiency score
Start with excel file 41467_2021_23576_MOESM4_ESM.xlsx from the CRISPRon paper in the Supplementary Data 1 section. In this file, the authors released the gRNA (20-mer) sequences and corresponding total_indel_eff scores. We specifically used the data from the sheet named spCas9_eff_D10+dox. We use the total_indel_eff column as the efficiency score.
Training and validation data
Given the 20-mers and the crispron scores (same as the total_indel_eff scores) from earlier, complete the following steps to put together the training and validation data:
- Convert the sequences in the sheet “TRAP12K microarray oligos” into an .fa (fasta) file.
- Run the script
get_30mers_from_fa.py(from the CRISPRon GitHub repository) to obtain all possible 23-mers and 30-mers from the sequences obtained from Step 1. - Use the
CRISPRspec_CRISPRoff_pipeline.pyscript (from the CRISPRon GitHub repository) to obtain the binding energy for the 23-mers obtained from Step 2. For more details on how to run this script, check out the code released by the authors of the CRISPRon paper(check the scriptCRISPRon.sh). - At this point, we have 23-mers along with the corresponding binding energy scores, and 20-mers along with the corresponding CRISPRon scores. Additionally, we have the 30-mers from Step 2.
- Use the script
prepare_train_dev_data.py(from our released code) to create training and validation splits. Running this script will create two files:train.csvanddev.csv.
The data looks something like the following:
Model architecture for gRNA encoding
To encode the gRNA sequence, we used the DNABERT encoder. DNABERT was pre-trained on human genomic data, so it’s a good model to encode gRNA sequences. DNABERT tokenizes the nucleotide sequence into overlapping k-mers, and each k-mer serves as a word in the DNABERT model’s vocabulary. The gRNA sequence is broken into a sequence of k-mers, and then each k-mer is replaced by an embedding for the k-mer at the input layer. Otherwise, the architecture of DNABERT is similar to that of BERT. After we encode the gRNA, we use the representation of the [CLS] token as the final encoding of the gRNA sequence. To predict the efficiency score, we use an additional regression layer. The MSE loss will be the training objective. The following is a code snippet of the DNABertForSequenceClassification model:
Fine-tuning and prompting genomic LLMs
Fine-tuning all the parameters of a model is expensive because the pre-trained model becomes much larger. LoRA is an innovative technique developed to address the challenge of fine-tuning extremely large language models. LoRA offers a solution by suggesting that the pre-trained model’s weights remain fixed while introducing trainable layers (referred to as rank-decomposition matrices) within each transformer block. This approach significantly reduces the number of parameters that need to be trained and lowers the GPU memory requirements, because most model weights don’t require gradient computations.
Therefore, we adopted LoRA as a PEFT method on the DNABERT model. LoRA is implemented in the Hugging Face PEFT library. When using PEFT to train a model with LoRA, the hyperparameters of the low rank adaptation process and the way to wrap base transformers models can be defined as follows:
Hold-out evaluation performances
We use RMSE, MSE, and MAE as evaluation metrics, and we tested with rank 8 and 16. Furthermore, we implemented a simple fine-tuning method, which is simply adding several dense layers after the DNABERT embeddings. The following table summarizes the results.
| Method | RMSE | MSE | MAE |
| LoRA (rank = 8) | 11.933 | 142.397 | 7.014 |
| LoRA (rank = 16) | 13.039 | 170.01 | 7.157 |
| One dense layer | 15.435 | 238.265 | 9.351 |
| Three dense layer | 15.435 | 238.241 | 9.505 |
| CRISPRon | 11.788 | 138.971 | 7.134 |
When rank=8, we have 296,450 trainable parameters, which is about 33% trainable of the whole. The performance metrics are “rmse”: 11.933, “mse”: 142.397, “mae”: 7.014.
When rank=16, we have 591,362 trainable parameters, which is about 66% trainable of the whole. The performance metrics are “rmse”: 13.039, “mse”: 170.010, “mae”: 7.157. There might have some overfitting issue here under this setting.
We also compare what happens when adding a few dense layers:
- After adding one dense layer, we have “rmse”: 15.435, “mse”: 238.265, “mae”: 9.351
- After adding three dense layers, we have “rmse”: 15.435, “mse”: 238.241, “mae”: 9.505
Lastly, we compare with the existing CRISPRon method. CRISPRon is a CNN based deep learning model. The performance metrics are “rmse”: 11.788, “mse”: 138.971, “mae”: 7.134.
As expected, LoRA is doing much better than simply adding a few dense layers. Although the performance of LoRA is a bit worse than CRISPRon, with thorough hyperparameter search, it is likely to outperform CRISPRon.
When using SageMaker notebooks, you have the flexibility to save the work and data produced during the training, turn off the instance, and turn it back on when you’re ready to continue the work, without losing any artifacts. Turning off the instance will keep you from incurring costs on compute you’re not using. We highly recommend only turning it on when you’re actively using it.
Conclusion
In this post, we showed how to use PEFT methods for fine-tuning DNA language models using SageMaker. We focused on predicting efficiency of CRISPR-Cas9 RNA sequences for their impact in current gene-editing technologies. We also provided code that can help you jumpstart your biology applications in AWS.
To learn more about the healthcare and life science space, refer to Run AlphaFold v2.0 on Amazon EC2 or fine-tuning Fine-tune and deploy the ProtBERT model for protein classification using Amazon SageMaker.
About the Authors
 Siddharth Varia is an applied scientist in AWS Bedrock. He is broadly interested in natural language processing and has contributed to AWS products such as Amazon Comprehend. Outside of work, he enjoys exploring new places and reading. He got interested in this project after reading the book The Code Breaker.
Siddharth Varia is an applied scientist in AWS Bedrock. He is broadly interested in natural language processing and has contributed to AWS products such as Amazon Comprehend. Outside of work, he enjoys exploring new places and reading. He got interested in this project after reading the book The Code Breaker.
 Yudi Zhang is an Applied Scientist at AWS marketing. Her research interests are in the area of graph neural networks, natural language processing, and statistics.
Yudi Zhang is an Applied Scientist at AWS marketing. Her research interests are in the area of graph neural networks, natural language processing, and statistics.
 Erika Pelaez Coyotl is a Sr Applied Scientist in Amazon Bedrock, where she’s currently helping develop the Amazon Titan large language model. Her background is in biomedical science, and she has helped several customers develop ML models in this vertical.
Erika Pelaez Coyotl is a Sr Applied Scientist in Amazon Bedrock, where she’s currently helping develop the Amazon Titan large language model. Her background is in biomedical science, and she has helped several customers develop ML models in this vertical.
 Zichen Wang is a Sr Applied Scientist in AWS AI Research & Education. He is interested in researching graph neural networks and applying AI to accelerate scientific discovery, specifically on molecules and simulations.
Zichen Wang is a Sr Applied Scientist in AWS AI Research & Education. He is interested in researching graph neural networks and applying AI to accelerate scientific discovery, specifically on molecules and simulations.
 Rishita Anubhai is a Principal Applied Scientist in Amazon Bedrock. She has deep expertise in natural language processing and has contributed to AWS projects like Amazon Comprehend, Machine Learning Solutions Lab, and development of Amazon Titan models. She’s keenly interested in using machine learning research, specifically deep learning, to create tangible impact.
Rishita Anubhai is a Principal Applied Scientist in Amazon Bedrock. She has deep expertise in natural language processing and has contributed to AWS projects like Amazon Comprehend, Machine Learning Solutions Lab, and development of Amazon Titan models. She’s keenly interested in using machine learning research, specifically deep learning, to create tangible impact.